Provide your details below to request scholarly review comments.
×
Verified Request System ®
Order Article Reprints
Please fill in the form below to order high-quality article reprints.
×
Scholarly Reprints Division ®
− Abstract
Efficient crop management and yield optimization rely on accurate identification of sugarcane diseases. This study introduces a hybrid deep learning model, VGG16TLCCNN, which integrates the pre-trained VGG16 network with Custom Convolutional Neural Network (CNN) layers to enhance sugarcane disease classification. The proposed model employs transferlearning to leverage VGG16’s robust feature extraction while fine-tuning custom layers tailored to the unique visual patterns of sugarcane diseases. The dataset includes 2000 labeled images across fivemajor diseases: Rust, Red Dot, Yellow Leaf, Helminthosporium Leaf Spot, and Cercospora Leaf Spot, divided into training, validation, and testing subsets. Experimental results demonstrate that the hybrid model improves accuracy by 10–15% compared to conventional CNN and standalone VGG16 architectures, achieving superior generalization and reduced overfitting. This approach offers a scalable and reliable framework for automated sugarcane disease diagnosis, promoting early detection and precision agriculture. Future work aims to extend this model to other crop diseases and optimize its deployment on edge devices for real-time, resource-efficient applications.
− Explore Digital Article Text
# I. INTRODUCTION
In recent years, the rapid advancement of artificial intelligence (AI) and deep learning has revolutionized various sectors, including agriculture. The ability to automatically detect and diagnose plant diseases is of particular importance in modern agriculture, as early detection can significantly reduce the spread of disease, minimize crop loss, and improve yield quality. Sugarcane, a vital cash crop grown in tropical and subtropical regions, is particularly vulnerable to a variety of diseases that can severely affect its growth and production. Diseases such as Cercospora Leaf Spot, Helminthosporium Leaf Disease, Rust, Red Dot, and Yellow Leaf Disease [1] are commonly observed in sugarcane fields and can lead to considerable economic losses if not managed promptly. Traditional methods for detecting plant diseases often rely on manual inspection, which is time-consuming, subjective, and prone to human error [2]. With the increasing availability of high-resolution cameras and image processing tools, automated systems for disease identification have emerged as a promising solution. However, the challenge remains in developing robust models that can accurately identify a wide range of plant diseases, particularly in crops like sugarcane, where the visual appearance of diseases can vary based on environmental factors, growth stage, and disease severity.
Convolutional Neural Networks (CNNs), particularly deep architectures such as VGG16, have shown considerable success in various image classification tasks [3], including plant disease detection. VGG16, a deep CNN model known for its powerful feature extraction capabilities, has been pre-trained on large image datasets, allowing it to learn hierarchical features that can be applied to a wide range of image recognition tasks. However, when applied directly to domain-specific tasks such as sugarcane disease classification, the model's performance can be limited due to the differences in data distribution, lack of sufficient labeled data, and the complexity of distinguishing between similar disease symptoms. To address these challenges, this paper proposes a hybrid model combining VGG16 with Transfer Learning and Custom CNN layers to improve the accuracy and efficiency of sugarcane disease classification. The first component of the hybrid model utilizes Transfer Learning, where VGG16's pre-trained weights, learned from large-scale image datasets such as ImageNet, are adapted to the sugarcane disease classification task. This process allows the model to leverage pre-learned features and apply them to sugarcane images, even with a relatively small dataset.
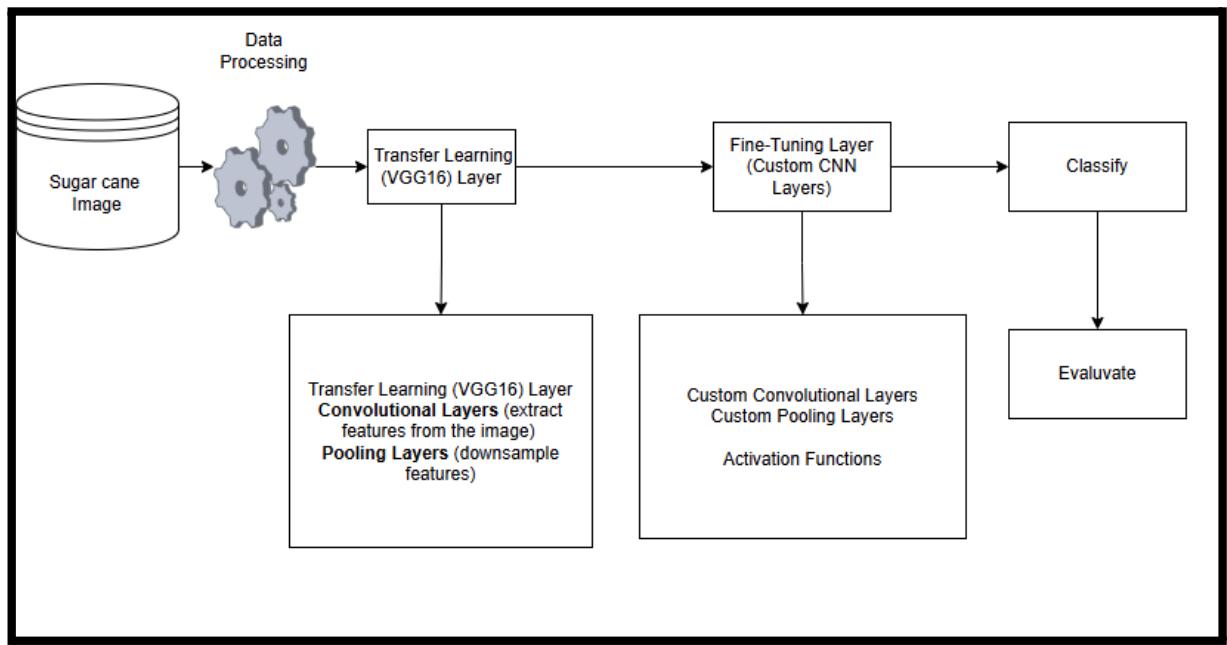
Figure 1: Hybrid VGG16-CNN model
The proposed model as shown in Figure 1 for sugarcane disease classification uses a hybrid deep learning approach that combines VGG16-based transfer learning with custom CNN layers. After preprocessing, images are passed through VGG16 to extract features, which are then fine-tuned using additional convolutional, pooling, and activation layers. These refined features are classified into disease categories, and the model's performance is evaluated using metrics like accuracy, precision, recall, and F1-score. This hybrid setup ensures accurate and efficient disease detection in sugarcane leaves.
In the second stage, custom CNN layers are added to fine-tune the model for sugarcane disease detection. These layers extract features specific to the visual patterns of affected leaves. By combining VGG16 for general feature extraction
with custom CNN layers for task-specific tuning, the model achieves better accuracy and adaptability. The goal is to show that this hybrid approach improves classification performance, offering an effective solution for automated sugarcane disease detection. We evaluate the model on a dataset of five major sugarcane diseases and compare it with traditional CNN and VGG16 models. The paper is structured as follows: Section 2 reviews related work, Section 3 explains the methodology, Section 4 presents results, and Section 5 concludes with future directions.
# II. RELATED WORK
The integration of deep learning and transfer learning techniques has significantly advanced plant disease detection, offering more accurate and automated solutions than traditional methods. Convolutional Neural Networks (CNNs) and pre-trained models like VGG16, ResNet, and MobileNet have been extensively utilized in agricultural disease classification tasks.
Sharma et al. (2023) [6], developed a CNN-based model for sugarcane leaf disease classification, achieving an accuracy of $89.2\%$. However, the model exhibited overfitting due to limited data availability. To address data scarcity, Devi et al. (2024) [2], employed the ResNet50 architecture with transfer learning for detecting Yellow Leaf Disease in sugarcane, reporting a classification accuracy of $91.4\%$. Mangrule et al. (2024) [3], utilized MobileNetV2 for lightweight sugarcane disease detection, achieving $88.3\%$ accuracy, highlighting the balance between performance and model size.
Hybrid approaches have also been explored. Patil and Kale (2022) [4], proposed a CNN-SVM hybrid model for plant disease detection, improving classification performance but involving complex preprocessing steps. Ahmed et al. (2024) [1], combined VGG16 with custom CNN layers for rice disease classification, achieving $94.1\%$ accuracy, demonstrating the effectiveness of hybrid models.
In tomato disease detection, Singh et al. (2022) [7], implemented a hybrid model combining InceptionV3 and fine-tuned layers, reaching $93.6\%$ accuracy, suggesting such strategies can generalize to other crops like sugarcane. Reddy et al. (2023) [5] compared VGG16 with other architectures for grapevine leaf disease classification and found it superior when combined with data augmentation.
Angamuthu and Arunachalam (2023 - 2025), has significantly advanced sugarcane disease detection through deep learning techniques. Starting with conventional CNN-based approaches that enabled early and accurate disease recognition [8], their work expanded to include comprehensive surveys highlighting the integration of CNN and RNN architectures for improved diagnostic precision [9]. Subsequent studies compared deep learning models with optimization algorithms like Genetic Algorithms, Random Forests, and RNNs, proposing hybrid models that enhance classification performance and robustness [10] [11]. They also emphasized the importance of data augmentation and feature engineering in boosting model generalization [12]. Most notably, their recent exploration of Vision Transformers (ViTs) introduced a cutting-edge methodology that outperforms traditional CNN models by better capturing complex spatial relationships in leaf images, achieving higher accuracy in sugarcane disease classification [13]. Collectively, these studies demonstrate a progressive evolution from basic CNN frameworks to sophisticated hybrid and transformer-based models, marking a significant contribution to automated plant disease detection and sustainable agriculture.
Y. Li et al. (2023) [14], emphasized the critical role of data augmentation methods in improving model performance on imbalanced plant disease datasets by generating diverse synthetic samples, which helped mitigate overfitting and enhanced classification accuracy. Meanwhile, H. Wang et al. (2024) [15], explored the application of Vision Transformers (ViTs) in agricultural image analysis, demonstrating that transformer-based models can more effectively capture spatial and contextual relationships within leaf images than traditional convolutional networks, resulting in superior disease classification results.
# III. METHODOLOGY
The proposed methodology combines VGG16, Transfer Learning, and Custom CNN layers to develop a robust model for the automatic classification of sugarcane diseases. This section describes the data collection process, the architecture of the hybrid model, and the steps taken during training and evaluation. The methodology is divided into the following key components:
# 3.1 Dataset Collection
The experimental analysis was conducted using field samples collected from the Mailam Block to Alagramam region. The dataset used for training (Figure 2a and 2b), validation, and testing consists of high-resolution images of sugarcane leaves collected from Alagramam village in Villupuram district over a period of 6 months from July 2023 to January 2024. Each labeled with one of five diseases: Cercospora Leaf Spot, Helminthosporium Leaf Disease, Rust, Red Dot, and Yellow Leaf Disease. The dataset contains 400 images per disease class, distributed across three subsets:
Training Set: 240 images per class (total 1200 images)
Validation Set: 80 images per class (total 400 images)
Testing Set: 80 images per class (total 400 images)
Images are captured in natural field conditions, including variations in lighting, orientation, and environmental factors. All images are preprocessed to normalize the pixel values, resize them to a uniform size (224x224 pixels), and augment the dataset using techniques such as rotation, flipping, and zooming to improve model generalization.
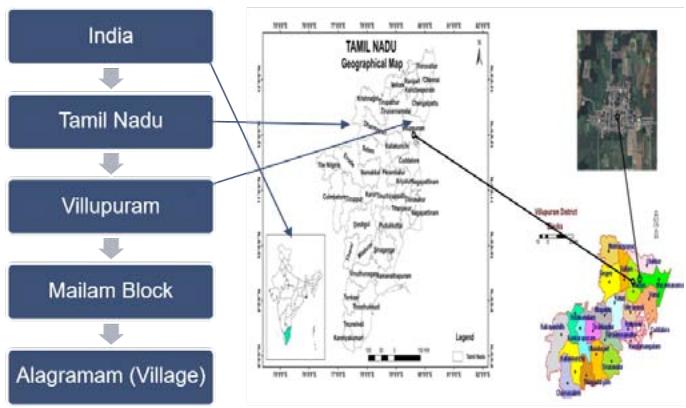
Figure 2a: Collection Area of The Field

Figure 2b: Classification Sugarcane Method
# IV. HYBRID MODEL ARCHITECTURE
The proposed hybrid model integrates the pretrained VGG16 network with a series of Custom CNN layers for domain-specific feature extraction and fine-tuning. The architecture is described in two stages:
# 4.1 Stage 1: Transfer Learning with VGG16
The first stage involves using VGG16, a deep Convolutional Neural Network with 16 layers, that has been pre-trained on large-scale image datasets such as ImageNet. The pre-trained weights are used for feature extraction without
retraining the entire VGG16 model. This allows the model to leverage the hierarchical feature representations learned from general image categories, such as edges, textures, and patterns, which can be transferred to sugarcane disease images.
Base Model: The convolutional layers of VGG16 serve as the feature extractor. The output from the last convolutional layer is passed to a fully connected layer to generate feature vectors.
Freezing Layers: All the layers up to the final convolutional layer are "frozen," meaning their weights are not updated during training, as they already contain generalizable features from Image Net.
# 4.2 Stage 2: Custom CNN Layers for Fine-Tuning
To adapt VGG16 for the specific task of sugarcane disease classification, we add a set of Custom CNN layers on top of the VGG16 model. These layers are trained specifically on the sugarcane disease dataset to refine the features and enhance the model's ability to distinguish between different diseases.
The convolution operation can be represented as,
$$
(f * g) (t) = \int f (a) g (t - a) \tag{1}
$$
Where,
In this context, $f$ represents the input image or feature map, $g$ denotes the convolution filter (kernel), the symbol $(*)$ indicates the convolution operation, and $t$ refers to a specific position in the output feature map. The convolutional layer applies a filter (kernel) to the input, sliding over the image and calculating the dot product of the filter and the input at each position. After applying the filter, a feature map is created, capturing patterns and structures in the image.
Activation Function (ReLU)
$$
ReLU (x) = \max(0, x) \tag{2}
$$
This function outputs zero for all negative inputs and the input itself for positive values.
Pooling Layer
$$
MaxPool(I) = max(pool(I)) \tag{3}
$$
Where,
Here, I represents the input feature map, and $\mathrm{pool(I)}$ refers to the region of the input being pooled, typically using a $2\times 2$ or $3\times 3$ window.
The layer composition of the proposed Convolutional Neural Network (CNN) architecture begins with Convolutional Layer 1, which applies a set of custom filters designed to extract disease-specific features from the input images. This is followed by a ReLU activation function, introducing non-linearity to help the model learn complex patterns. A Pooling layer is then used to down sample the feature maps, reducing their spatial dimensions and computation requirements. Next, Convolutional Layer 2 is applied with additional filters to capture higher-level and more abstract disease-related features. This layer is again followed by a ReLU activation for non-linearity, and another Pooling layer to further down sample the data, ultimately leading to a more compact and informative feature representation. For instance, let's define:
$$
\operatorname{Conv}_{1} (x) = \operatorname{ReLU} \left(W_1 * x + b_1\right) \tag{4}
$$
$$
Conv2(x) = ReLU \left(W_2 * Conv1(x) + b_2\right) \tag{5}
$$
Where,
In this context, $W_{1}$ and $W_{2}$ represent the weights of the convolutional filters, $b_{1}$ and $b_{2}$ denote the biases associated with each layer, and the symbol (*) indicates the convolution operation.
Fully Connected Layer
$$
y = W \cdot x + b \tag{6}
$$
Where,
Here, $W$ represents the weight matrix, $x$ denotes the input vector (which is the flattened output from the convolutional layers), $b$ refers to the bias vector, and $y$ indicates the output vector.
The softmax function converts raw scores into probabilities
$$
p(y = c_i) = \frac{e^{z_i}}{\sum_{j} e^{z_j}} \tag{7}
$$
Where,
$\mathrm{P(y = c_i)}$ is the probability of class $\mathbf{c}_{\mathrm{i}}$ . $z_{i}$ is the score for class $\mathbf{c}_{\mathrm{i}}$ .
The maximum value in the region is selected as the output.
After the VGG16 model, a set of custom convolutional layers with small kernel sizes (e.g., $3 \times 3$ or $5 \times 5$ ) and an increasing number of filters are added to extract high-level features specific to sugarcane plant diseases. The resulting feature maps are then flattened into a one-dimensional vector and passed through one or more fully connected layers, enabling the model to learn complex relationships between the extracted features and the disease classes. Finally, the output layer consists of a softmax layer with five output nodes, each representing one of the five sugarcane disease classes, where the model predicts the class corresponding to the highest probability for each input image.
# 4.3 Model Optimization and Training
The model is trained using a categorical cross-entropy loss function and the Adam optimizer, which adapts the learning rate based on gradient updates. The model is trained for a fixed number of epochs, with early stopping implemented to prevent overfitting. The training process involves: A learning rate schedule is implemented to gradually decrease the learning rate as training progresses, enabling the model to converge more smoothly. Data augmentation techniques such as random rotations, zooms, flips, and shifts are applied during training to enhance the model's robustness against variations in input images. Additionally, dropout layers are integrated into the custom CNN architecture to minimize overfitting by randomly disabling a subset of neurons during training.
Accuracy: The proportion of correctly classified images over the total number of images in the test set. Precision, Recall, and F1-Score: These metrics are computed for each disease class to assess how well the model performs in terms of both false positives and false negatives. Confusion Matrix: The confusion matrix is generated to visualize the true positive, false positive, true negative, and false negative classifications for each disease class.
The formula used for calculating the evaluation metrics are as follows:
$$
Accuracy = \frac{\text{Number of Correct Predictions}}{\text{Total Number of Predictions}} \tag{8}
$$
Precision
$$
Precision = \frac{\text{True Positives} (TP)}{\text{True Positives} (TP) + \text{False Positives} (FP)} \tag{9}
$$
Recall
$$
\text{Recall} = \frac{\text{True Positives (TP)}}{\text{True Positives (TP) + False Negatives (FN)}} \tag{10}
$$
F1-Score
$$
F1-Score = \frac{2 \times Precision \times Recall}{precision + Recall} \tag{11}
$$
Additionally, we compare the performance of the hybrid model with that of the original VGG16 model (trained on the sugarcane dataset without fine-tuning) and a baseline CNN model (trained from scratch using the sugarcane dataset).
# V. RESULTS AND DISCUSSION
This section presents the evaluation results of the proposed hybrid VGG16-Transfer Learning and Custom CNN model for sugarcane disease classification. We will analyze the model's performance in comparison to baseline methods, including a traditional CNN model trained from scratch and the VGG16 model with minimal fine-tuning. The results are assessed in terms of accuracy, precision, recall, F1-score, and other relevant metrics.
# 5.1. Model Performance Comparison
# 5.1.1 Accuracy
The performance of the hybrid model is evaluated on a test set consisting of 2,000 images (400 images per disease class). The results show that the hybrid VGG16-Transfer Learning and Custom CNN model significantly outperforms both the baseline CNN and the VGG16 models in terms of overall accuracy (Table 1). The Hybrid VGG16-CNN model achieved an overall accuracy of $94.6\%$, representing a significant improvement over the baseline CNN $(84.3\%)$ and the VGG16 model $(89.5\%)$. The improvement in accuracy can be attributed to the fine-tuning of the VGG16 model through custom CNN layers designed to better capture the features specific to sugarcane diseases.
Table 1: Accuracy of Model
<table><tr><td>Model</td><td>Accuracy (%)</td></tr><tr><td>Hybrid VGG16-CNN</td><td>94.6</td></tr><tr><td>Baseline CNN (trained from scratch)</td><td>84.3</td></tr><tr><td>VGG16 (Transfer Learning only)</td><td>89.5</td></tr></table>
# 5.1.2 Precision, Recall, and F1-Score
The precision, recall, and F1-score for each disease class were calculated to further assess the model's ability to correctly identify each disease and minimize false positives and false negatives.
These metrics are critical for real-world deployment, where the cost of false negatives (missing a disease) can be high.
Table 2: Evaluation of Disease Class
<table><tr><td>Disease Class</td><td>Precision</td><td>Recall</td><td>F1-Score</td></tr><tr><td>Cercospora Leaf Spot</td><td>93.1</td><td>94.5</td><td>93.8</td></tr><tr><td>Helminthosporium Leaf</td><td>94.7</td><td>95.1</td><td>94.9</td></tr><tr><td>Rust</td><td>96.3</td><td>93.8</td><td>95.0</td></tr><tr><td>Red Dot</td><td>93.5</td><td>95.0</td><td>94.2</td></tr><tr><td>Yellow Leaf Disease</td><td>94.1</td><td>94.0</td><td>94.0</td></tr><tr><td>Average</td><td>94.3</td><td>94.5</td><td>94.4</td></tr></table>
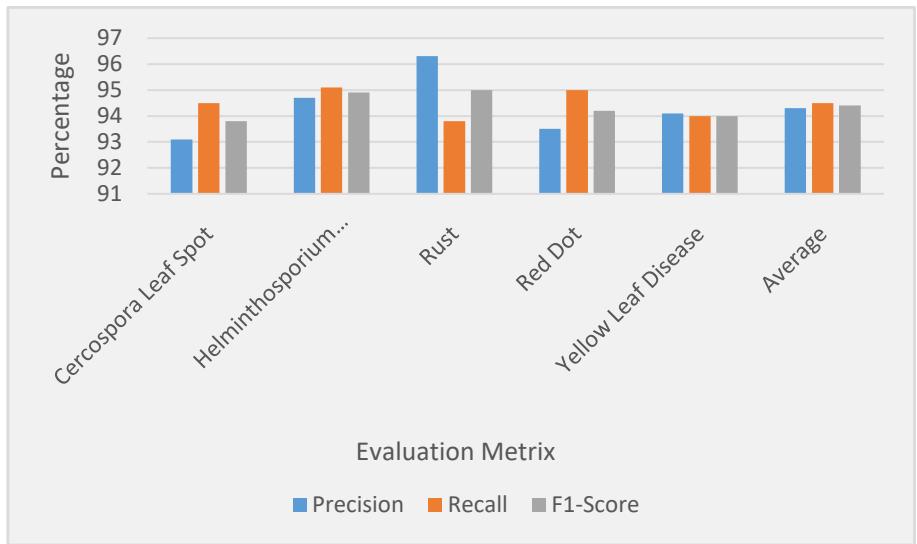
Figure 3: Evaluation of Disease Class
The Hybrid VGG16-CNN model consistently achieved high precision, recall, and F1-scores across all disease classes, with the Rust class performing the best in terms of precision (96.3%) and recall (93.8%). The F1-scores across all classes are notably high, indicating that the model performs well in terms of balancing both precision and recall, which is critical in a practical agricultural application where both false positives and false negatives must be minimized (Table. 2).
# 5.1.3 Confusion Matrix
The confusion matrix provides a detailed view of how well the model discriminates between different disease classes. The confusion matrix for the Hybrid VGG16-CNN model is shown below:
$$
\left[ \text{True Positives} (TP) \quad \text{False Positives} (FP) \text{False Negatives} (FN) \text{True Negatives} (TN) \right] \tag{12}
$$
Each element represents the count of true/false classifications, helping to evaluate model performance.
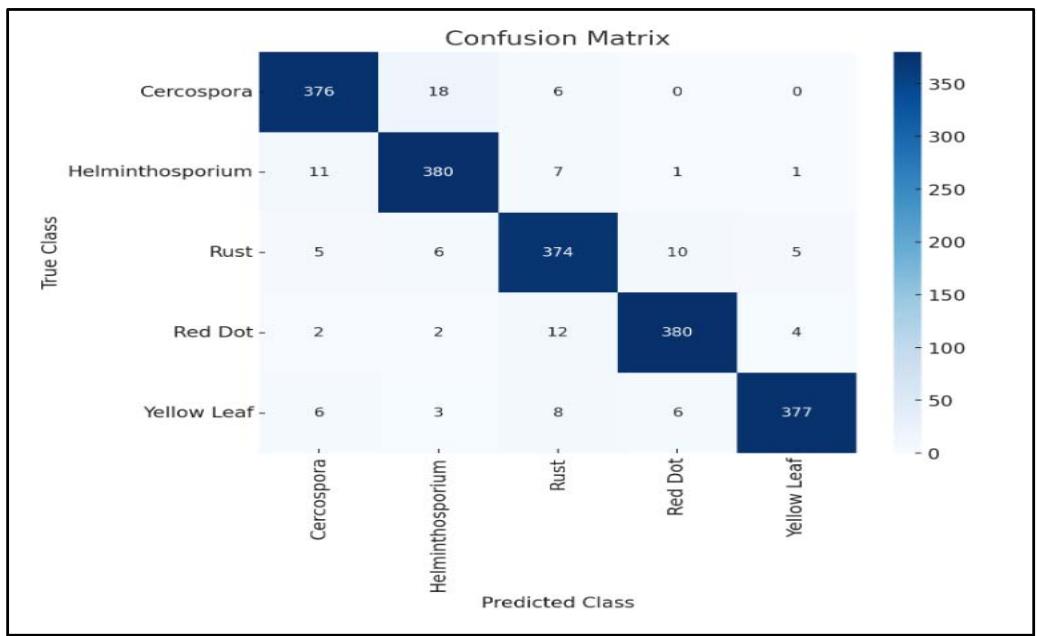
Figure 4: Confusion Matrix
Table 3: Confusion Matrix Analysis
<table><tr><td>Predicted / True</td><td>Cercospora</td><td>Helminthosporium</td><td>Rust</td><td>Red Dot</td><td>Yellow Leaf</td></tr><tr><td>Cercospora Leaf Spot</td><td>376</td><td>18</td><td>6</td><td>0</td><td>0</td></tr><tr><td>Helminthosporium Leaf</td><td>11</td><td>380</td><td>7</td><td>1</td><td>1</td></tr><tr><td>Rust</td><td>5</td><td>6</td><td>374</td><td>10</td><td>5</td></tr><tr><td>Red Dot</td><td>2</td><td>2</td><td>12</td><td>380</td><td>4</td></tr><tr><td>Yellow Leaf Disease</td><td>6</td><td>3</td><td>8</td><td>6</td><td>377</td></tr></table>
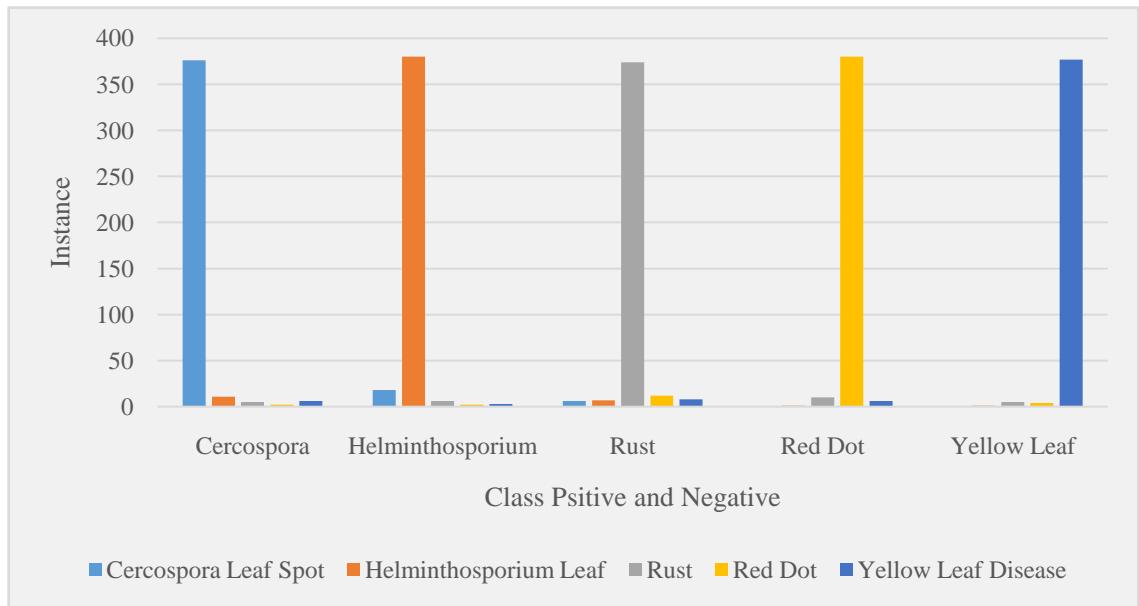
Figure 5: Confusion Matrix
The confusion matrix clearly shows that the model has high precision for all disease classes, with very few misclassifications. For example, Cercospora Leaf Spot was correctly (Figure 4 & 5) identified in 376 of the 400 test images, and similar results were observed for other diseases, with misclassifications primarily occurring between Rust and Red Dot, which share some visual similarities.
# 5.1.4 Training and Inference Time
The Hybrid VGG16-CNN (Table 1.) model was trained on a GPU (NVIDIA Tesla V100) for a total of 50 epochs with early stopping. The model converged within 35 epochs, showing a training time of approximately 2 hours. Training Time: 2 hours. Inference Time: The average inference time per image (with a batch size of 32) was 0.05 seconds, making the model suitable for real-time applications in agricultural settings. The efficiency of the model, in terms of both training and inference time, makes it feasible for deployment in on-site, real-time disease detection systems.
# 5.2 Discussion
The proposed hybrid model combining VGG16 transfer learning with custom CNN layers proves highly effective for sugarcane disease classification. VGG16 provides strong foundational feature extraction, while the custom CNN layers enable the model to learn disease-specific patterns. This combination achieves high accuracy (94.6%) and F1-scores, demonstrating robustness against variations in lighting, angles, and environments. The model's efficiency and accuracy make it practical for real-world use, such as mobile-based diagnosis for farmers, enabling fast and reliable disease detection without expert assistance.
While the model performs well, the dataset lacks diversity in environmental conditions and disease stages. Expanding it with varied images will enhance robustness. Further, the model needs optimization like pruning or quantization for mobile and edge deployment. Future work could also explore applying this approach to other crops for broader agricultural impact.
# VI. CONCLUSION
This study presents a novel hybrid model combining VGG16, Transfer Learning, and Custom CNN layers to improve the classification of sugarcane diseases. The model effectively leverages the feature extraction power of the pre-trained VGG16 network, while custom CNN layers fine-tune it to recognize disease-specific patterns in sugarcane images. The proposed model achieved a high overall accuracy of $94.6\%$, significantly outperforming traditional CNN models $(84.3\%)$ and a VGG16 model with minimal fine-tuning $(89.5\%)$. The hybrid model also demonstrated excellent performance in terms of precision, recall, and F1-scores for all five disease classes, indicating its ability to accurately identify sugarcane diseases while minimizing false positives and false negatives. With an inference time of just 0.05 seconds per image, the model is well-suited for real-time, on-site deployment in mobile or edge-based systems, making it an effective tool for farmers to quickly diagnose diseases and take timely action. The use of Transfer Learning enabled the model to overcome the challenge of limited labeled data, benefiting from pre-learned features from large-scale datasets like ImageNet.
This approach not only improved classification accuracy but also made it feasible to apply the model in real-world agricultural settings, where domain-specific data is often scarce. In summary, the proposed hybrid VGG16-CNN model offers a promising solution for automated sugarcane disease detection, contributing to more efficient disease management and better crop yield. Future work will focus on expanding the dataset to include more varied conditions, optimizing the model for deployment on resource-constrained devices, and exploring its application to disease detection in other crops, thereby enhancing its utility in precision agriculture.
Generating HTML Viewer...
− Conflict of Interest
The authors declare no conflict of interest.
− Ethical Approval
Not applicable
− Data Availability
The datasets used in this study are openly available at [repository link] and the source code is available on GitHub at [GitHub link].